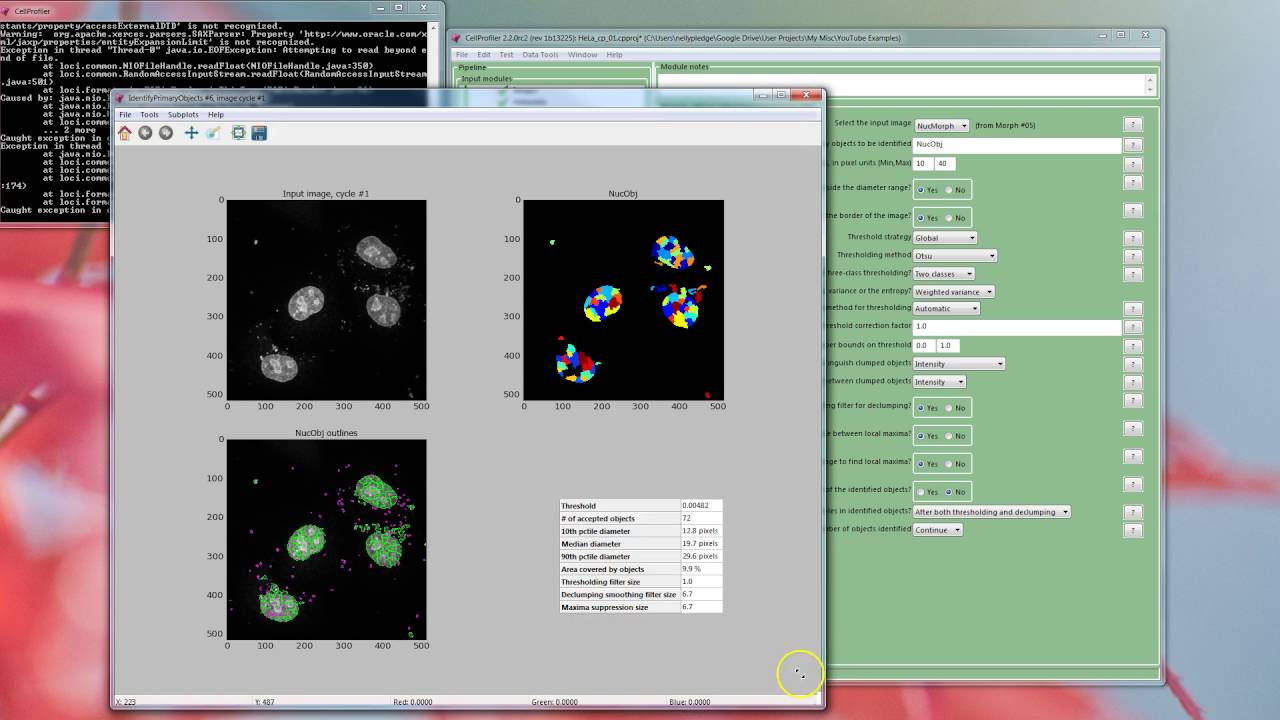 
Two cell types have been stained that are of particular interest in Hodgkin’s Lymphoma: Reed-Sternberg cells (aka RS in green), and tumor-associated macrophages (aka TAM in red). The nuclei of all cells are stained (blue). Consider the image of a representative tissue slice (below), which reveals a field of view with a high cell density. In collaboration with the Margaret Shipp and Scott Rodig labs, we developed a pipeline in CellProfiler that addresses unique challenges presented by imaging tissue slices. Ultimately the aim is to define various configurations of this interaction that are predictive of patient outcome or the likelihood of success for a given treatment, such as immunotherapy. The ability to precisely measure this relationship will give a deeper understanding of the progression of cancer and might yield new insight into when and how the immune system is involved. Quantifying the cell locations provides the ability to gauge the degree to which the cancer has invaded a tissue and how the immune system is interacting with the leading edge of a tumor. In lymph node sections, cells have representatives from the immune system, epithelial tissue, connective tissue, and cancer. Quantifying the spatial relationship among cells in the crowded environment of a tissue requires reliable segmentation of several cell types. The strength of the inflammatory response has been linked to the prognosis of certain cancers such as lymphoma. Interactions among cells within a tissue are crucial to understanding the role of the inflammation that is triggered by the invasion of cancerous cells. This is an abstract that was submitted for the 2016 ARVO Annual Meeting, held in Seattle, Wash., May 1-5, 2016.Imaging tissue slices provides a wealth of data about the spatial composition and number of the various cell types that make up a tissue. This approach would work well with many other histology or EM image sets. Our automated tool has become an integral part of our RPE sheet and cell analysis system. This automated method proved useful for the analysis of RPE cells in flat mounts with greater accuracy, more efficient sampling of data points, and less human bias and errors. The risk of human errors, heavy time commitment, and sampling bias are just some of the drawbacks of manual implementations. Manual analysis of image files, such as images of retinal pigment epithelium (RPE) sheet, is difficult and in many cases, quantitative manual analysis of cell shapes is impossible. This method allows almost completely automated processing once images were collected, ensuring collection of more quantitative and unbiased data. With this method, coverage of the whole RPE sheet was increased from about 20% to 50%. While the manual selection method took 1.5-2.5 hours for each eye, our automated counterpart reduced completion time to a few seconds. Further, the scripts numbered the boxes, and collected x,y coordinates automatically, and organized them for input into CellProfiler version 2.1.1, which then performed downstream analysis automatically. We implemented an automated method based on statistical analysis using k-means clustering of the tissue patterns to automatically generate and select artifact-free boxes from the RPE sheet. CellProfiler software was used to perform morphometric analysis of cell shape, area, number of neighboring cells, and 20 other metrics.Ī major bottleneck in processing images is to select parts of the RPE sheet that lacked dissection, processing, and staining artifacts. Images were processed using MATLAB scripts and an automated “Cutbox selector”. The RPE sheet was prepared and imaged as before (Boatright et al. RPE flat mounts were compared from two groups of mice, rd10 and C57BL/6J at different ages (P30, P45, P60 and P100). Bulk analysis of such data sets helps us to measure individual cell morphometry, and to determine texture and spatial point properties of RPE sheets that reflect cell-cell and cell-environment interactions and cell organization in normal and diseased eyes. This study aimed to develop an automated method for analyzing large series of retinal pigment epithelium (RPE) cells in flat mounts.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
